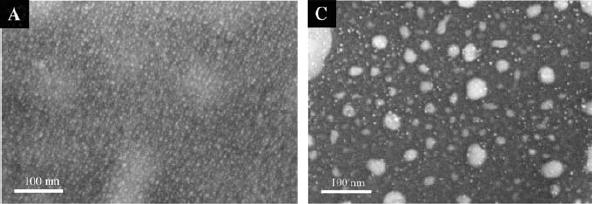
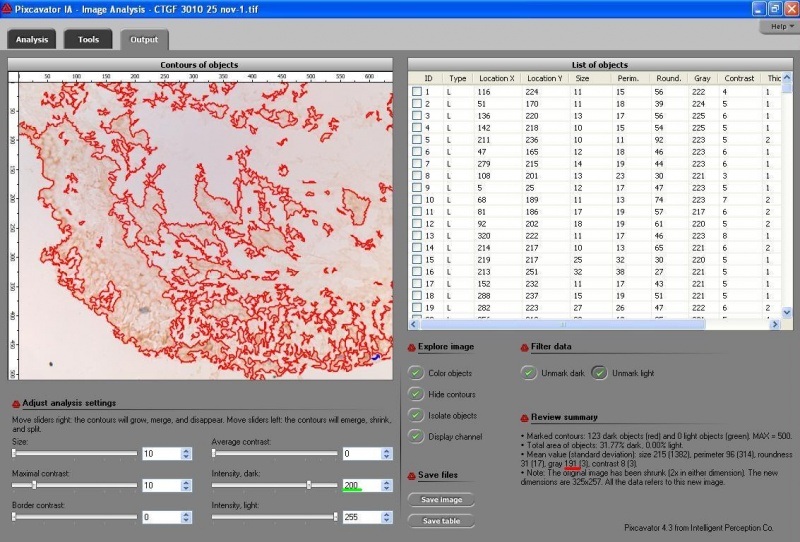
Pixcavator 5.0 released
These are the new features in version 5.0.
- Your choice of settings in the Output tab (the position of the sliders) is preserved when you load a new image to analyze.
- Your choice of color channels in the Analysis tab is preserved when you load a new image to analyze. With these two the user can apply the same settings to a sequence of images if they are similar in nature. So, we get as close as possible to bulk processing without actually creating this complex feature.
- Luminosity is a new color channel that you can choose. It is computed as a combination of the red, green, and blue values: 0.299*R + 0.587*G + 0.114*B. There are four channels now.
- “Display channel” is a new option in the Analysis tab (just like the one in the Output tab). If you have chosen to shrink the image, the shrunken version is shown. This way you can preview all channels and decide which is the best - before committing to time consuming analysis.
- The “Help” menu provides now the links to the help pages of this wiki. The user’s guide and the license are still provided with the program; they are to be found in the “Pixcavator” folder on your hard disk.
- The actual processing time is shown when it’s done, and a beep is produced - but only if processing has taken more than 5 seconds.
- Up to 2000 contours are now shown on the image and their statistics is also displayed. When there are more than 2000 contours, neither is shown.
- A few bugs have been fixed, some remain.
Download here.
Digital discoveries
- Casinos Not On Gamstop
- Non Gamstop Casinos
- Casino Not On Gamstop
- Casino Not On Gamstop
- Non Gamstop Casinos UK
- Casino Sites Not On Gamstop
- Siti Non Aams
- Casino Online Non Aams
- Non Gamstop Casinos UK
- UK Casino Not On Gamstop
- Non Gamstop Casino UK
- UK Casinos Not On Gamstop
- UK Casino Not On Gamstop
- Non Gamstop Casino UK
- Non Gamstop Casinos
- Non Gamstop Casino Sites UK
- Best Non Gamstop Casinos
- Casino Sites Not On Gamstop
- Casino En Ligne Fiable
- UK Online Casinos Not On Gamstop
- Online Betting Sites UK
- Meilleur Site Casino En Ligne
- Migliori Casino Non Aams
- Best Non Gamstop Casino
- Crypto Casinos
- Casino En Ligne Belgique Liste
- Meilleur Site Casino En Ligne Belgique
- Bookmaker Non Aams
- カジノ ライブ
- онлайн казино с хорошей отдачей
- スマホ カジノ 稼ぐ
- ブック メーカー オッズ
- Top 3 Nhà Cái Uy Tín Nhất
- Trang Web Cá độ Bóng đá Của Việt Nam
- Casino En Ligne Avis
- Casino En Ligne France
- Casino En Ligne
- 꽁머니 토토
- Casino Online Non Aams
- Meilleur Casino En Ligne