To speak at Bio-Image Informatics Workshop
Dr. Ash Pahwa and I will give a presentation at Workshop on Bio-Image Informatics: Biological Imaging, Computer Vision and Data Mining, 2008, Center for Bio-Image Informatics, UCSB, Santa Barbara, CA, USA, January 17-18, 2008. The abstract is below.
A Topological Approach to Cell Counting
Peter Saveliev (Marshall University, Huntington, WV) and Ash Pahwa (Mayachitra Inc., Santa Barbara, CA)
Cell counting and identification is a common task in biology and pathology. To automate this task one has to approach it as an image segmentation problem. Many researchers have solved this problem following various strategies. The approach we propose relies on topology, which is the science of continuity and connectedness that studies spatial relations within the image. An image pixel is defined to have 4 vertices (corners), 4 edges, and one face. Algebraic topology uses algebraic operations with these objects to count the number of completed cycles – circular sequences of edges. The completion of a cycle indicates the presence of a cell. In the case of gray scale, our strategy for counting cells is to count dark objects with light background and light objects with dark background. The types of images our algorithm is most suitable for are those that represent something 2-dimensional (rather than 2D images of 3D objects) such as images of cellular tissue or blood cells under a microscope.
The topological nature of the algorithm makes it especially suitable for cell counting. First, the count of cells is independent of their locations. Second, the measurements of cells are independent of their orientations with respect to the image grid. Third, the algorithm captures cells and other features regardless of their sizes, shapes, and locations with no deformation, smoothing, blurring or approximation.
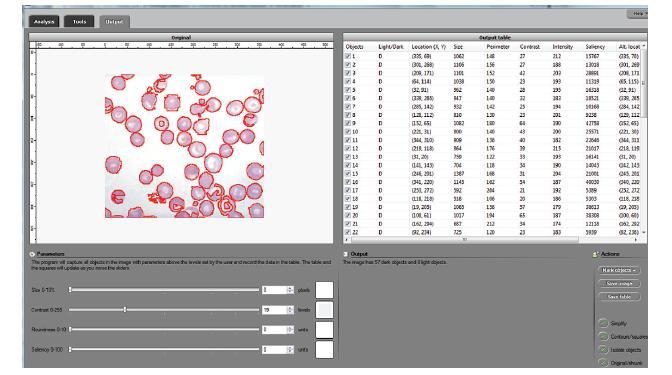
We developed a software suite called cellAnalyst with the following output: the image with cells’ contours captured and a spreadsheet with cells’ locations and characteristics such as area, perimeter, intensity, and contrast. The processing starts with an automatic analysis of the image that produces a graph that contains complete data about the image. The user proceeds in a semiautomatic mode to interactively visualize various segmentations. By moving sliders corresponding to cells’ characteristics the user instantly changes the cells’ boundaries and can choose the most appropriate segmentation. The output data is then updated in real time. The user can also exclude noise and irrelevant details from the analysis by simply clicking on them.
The analysis data has been verified using pathology and retinal images. The images are analyzed by manually counting, identifying, and measuring cells and the results are compared with the output of cellAnalyst. The matches have been reliable and repeatable.
Digital discoveries
- Casinos Not On Gamstop
- Non Gamstop Casinos
- Casino Not On Gamstop
- Casino Not On Gamstop
- Non Gamstop Casinos UK
- Casino Sites Not On Gamstop
- Siti Non Aams
- Casino Online Non Aams
- Non Gamstop Casinos UK
- UK Casino Not On Gamstop
- Non Gamstop Casino UK
- UK Casinos Not On Gamstop
- UK Casino Not On Gamstop
- Non Gamstop Casino UK
- Non Gamstop Casinos
- Non Gamstop Casino Sites UK
- Best Non Gamstop Casinos
- Casino Sites Not On Gamstop
- Casino En Ligne Fiable
- UK Online Casinos Not On Gamstop
- Online Betting Sites UK
- Meilleur Site Casino En Ligne
- Migliori Casino Non Aams
- Best Non Gamstop Casino
- Crypto Casinos
- Casino En Ligne Belgique Liste
- Meilleur Site Casino En Ligne Belgique
- Bookmaker Non Aams
- カジノ ライブ
- онлайн казино с хорошей отдачей
- スマホ カジノ 稼ぐ
- ブック メーカー オッズ
- Top 3 Nhà Cái Uy Tín Nhất
- Trang Web Cá độ Bóng đá Của Việt Nam
- Casino En Ligne Avis
- Casino En Ligne France
- Casino En Ligne
- 꽁머니 토토
- Casino Online Non Aams
- Meilleur Casino En Ligne












November 19th, 2007 at 7:49 pm
This is exciting. Is it available for use as yet? I am interested in looking at peripheral blood smear images.
November 19th, 2007 at 9:43 pm
Well, you can simply click “For researchers” above and try Pixcavator 2.3. It has served as a prototype for the cellAnalysist’s image analysis engine (CellAnalyst itself will also have database capabilities). CellAnalyst has been mostly put together but there is some work left. Also, before it becomes available we will have to do a lot more testing. Let me know if you have any questions, or if you have images you want me to try. — Peter