This site contains: mathematics courses and book; covers: image analysis, data analysis, and discrete modelling; provides: image analysis software. Created and run by Peter Saveliev.
Homology
From Intelligent Perception
The goal of this article is to provide an accessible overview of homology theory. To understand the challenge try this chapter from "A First (!) Course in Topology" by McCleary. One might instead start with Homology in 2D, Homology of cubical complexes, and Topological Features of Images or Homology as an equivalence relation.
1 Motivation from image analysis
The main principle of cell decomposition of images:
- The image is made of $k$-cells that are attached to each other along $(k-1)$-cells.
Here $k$ refers to the dimension of the cell.
In 2D we describe $0$- and $1$-cycles as circular sequences of edges. We were able to get away with this informality because of the peculiarity of the 2D case. First, the boundaries of objects can be traversed sequentially. Also, unlike in higher dimensions, there are no “intermediate” dimensions. For the topological features of dimensions between $0$ and the dimension of the image we have to use a more precise mathematical description.
In the $n$-dimensional case, the topological features we want captured are:
- "objects", blobs, or connected components, captured by $0$-cycles,
- holes or tunnels, captured by $1$-cycles, and
- voids or cavities, captured by $2$-cycles.
- etc.
The number of cycles of each kind is counted by the Betti numbers:
B0 B1 B2 homology Letter O 1 1 0 R, R, 0 Two letters O 2 2 0 R2,R2,0 Letter B 1 2 0 R1,R2,0 Donut 1 1 0 R1,R1,0 Tire 1 2 1 R1,R2,R1 Ball 1 0 1 R1,0,R1
2 The boundary operator
This is where topology becomes algebraic topology. The relations between $k$-cells and their boundaries as combinations of $(k-1)$-cells are recorded by mathematical formulas. For example, if $P$ is a pixel and $E_1,E_2,E_3,E_4$ are its edges then its boundary is $$DP = E_1+ E_2+ E_3+ E_4.$$ Here $DP$, or $D(P)$ if you prefer, is the boundary of $P$, so $D$ is the boundary operator.
Cells of all dimensions are easily recorded as lists of vertices:
- edge $E = (V_1, V_2)$ is the list of its end points,
- pixel $P = (V_1,V_2, V_3, V_4)$ is the list of its corners,
- voxel $X = (V_1, V_2, V_3, V_4, V_5, V_6, V_7, V_8)$ is the list of its corners,
- etc.
Suppose that both the pixel P and its edges and $E_1, E_2, E_3, E_4$ are oriented clockwise, then $$DP = E_1 + E_2 + E_3 + E_4.$$ But if pixel Q is oriented clockwise and its edges $G_1, G_2, G_3, G_4$ counterclockwise, then $$DQ = – G_1 – G_2 – G_3 – G_4.$$ Suppose now that P and Q are adjacent, say, Q lies to the right of P. Suppose edges $E_1, E_2, E_3, E_4$ and $G_1, G_2, G_3, G_4$ go around each of the pixels starting from the left low corner. Then $P$ and $Q$ share an edge, in fact, $E_3 = G_4$. Then, we compute algebraically the boundary of the combination of $P$ and $Q$: $$D(P + Q) = DP + DQ$$ $$= (E_1 + E_2 + E_3 + E_4) + (– G_1 – G_2 – G_3 – G_4) $$ $$= E_1 + E_2 + E_4 – G_1 – G_2 – G_3 $$ $$= E_1 + E_2 – G_3 – G_2 – G_1 + E_4.$$ Thus $E_1, E_2, -G_3, -G_2, -G_1, E_4$ go clockwise around $P + Q$, as expected.
Now, cells are combined into chains. For example, $P$ and $Q$ are $2$-chains consisting of a single $2$-cell each. Next, $DP$ and $DQ$ are $1$-chains as sums of four $1$-cells each. Thus, the boundary of a $k$-cell, such as $E, P$, or $Q$, or a $k$-chain, such as $P + Q$, is a $(k-1)$-chain. Here’s another example: $$DE = V_2 – V_1.$$ Moreover, both $DP$ and $DQ$ are “closed” $1$-chains in the sense that the $1$-cells form a circular curve. Such chains are called cycles. However, $DP$ and $DQ$ are trivial $1$-cycles in the sense that they are boundaries of $2$-chains, $P$ and $Q$. We are interested in nontrivial cycles such as $C$ and $C’$. They represent holes in the image.
Exercise 1. Compute $D(P – Q)$. Compare to $D(P + Q)$.
Exercise 2. Compute $DC, DC’, D(C + C’)$. What is $DK$ if $K$ is any cycle?
Exercise 3. Compute $DDP$. Draw conclusions. Answer
One can define homology with arbitrary coefficients, as long as it's a ring. Here we introduce homology for the first time with coefficients over $Z_2$. Keep in mind that integral homology above is the "best" homology -- just as base e is the best base of logarithms. In fact, it's even more so, see the Universal Coefficient Theorem.
3 Cycles and homology classes
Recall that cells of all dimensions are recorded as lists of vertices:
- edge $E = (V_1, V_2)$ is the list of its end points,
- pixel $P = (V_1,V_2, V_3, V_4)$ is the list of its corners,
- voxel $X = (V_1, V_2, V_3, V_4, V_5, V_6, V_7, V_8)$ is the list of its corners,
Now, $DP$ is a a cycle. A chain $C$ is easy to identify as a cycle because in this case $DC = 0$.
The 2D algorithm is extended to process nD cell complexes (simplicial, cubical, geometric, or CW-). In general, cell complexes are made of cells that can have an arbitrary number of faces of all dimensions and different shapes. It is assumed that a k-cell is homeomorphic to a k-ball. Thus, a $k$-cell is a $k$-dimensional polygon, possibly curvilinear, together with its orientation, i.e., the order of vertices. Further, $k$-cells are attached to each other only along their boundaries, which are made of $(k-1)$-cells.
Partition is now just a part of a more general topological decomposition of the image.
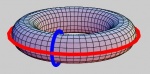
Consider the picture of the torus. The goal is to capture the tunnel that goes along the tire. Note, however, the tunnel is impossible to see if you are standing on the surface. The reason is that the only information you have is how these cells are attached to each other. The only way to detect the tunnel is by considering the cycles that go around it.
Now, the problem is that for just one tunnel there are infinitely many of such cycles. For example, every “meridian” goes around the tunnel, not just the blue one. The answer to this problem is to say they are all equivalent, or “homologous”, and they are counted as one. These cycles are indeed homologous (similar) as one can be slid to another along the surface, while the blue and the red are not. Algebraically, this works as follows.
A k-homology class in the image is represented by a chain of k-cells that can be recorded as an $M$-vector, where $M$ is the total number of $k$-cells in the frame. As an illustration, a component of this vector is $1$ if the corresponding cell is present, $-1$ if it present with the opposite orientation, $0$ if it is absent.
The $k$-boundary operator $D$ is an $M×N$ matrix where N, the number of rows, is the total number of $(k-1)$-cells in the image. Then the boundary of a chain $C$ is the matrix product $DC$. By means of $D$, it is easy to verify whether any two k-chains $A$ and $B$ are homologous: the difference between them is the boundary of a $(k+1)$-chain $T: A – B = DT$. The existence of $T$ is verified by the standard linear algebra techniques. In this case, $A$ and $B$ belong to the same $k$-homology class $H = [A] = [B]$.
Examples of such A and B are: two vertices in the same component of any cell complex ($k = 0$); two meridians of a torus, but not a meridian and the equator ($k = 1$); the inner surface and the outer surface of a thick sphere ($k = 2$).
In the presence of noise one has to consider persistent homology.
Consider also Homology software.
Digital discoveries
- Casinos Not On Gamstop
- Non Gamstop Casinos
- Casino Not On Gamstop
- Casino Not On Gamstop
- Non Gamstop Casinos UK
- Casino Sites Not On Gamstop
- Siti Non Aams
- Casino Online Non Aams
- Non Gamstop Casinos UK
- UK Casino Not On Gamstop
- Non Gamstop Casino UK
- UK Casinos Not On Gamstop
- UK Casino Not On Gamstop
- Non Gamstop Casino UK
- Non Gamstop Casinos
- Non Gamstop Casino Sites UK
- Best Non Gamstop Casinos
- Casino Sites Not On Gamstop
- Casino En Ligne Fiable
- UK Online Casinos Not On Gamstop
- Online Betting Sites UK
- Meilleur Site Casino En Ligne
- Migliori Casino Non Aams
- Best Non Gamstop Casino
- Crypto Casinos
- Casino En Ligne Belgique Liste
- Meilleur Site Casino En Ligne Belgique
- Bookmaker Non Aams
- カジノ ライブ
- онлайн казино с хорошей отдачей
- スマホ カジノ 稼ぐ
- ブック メーカー オッズ
- Top 3 Nhà Cái Uy Tín Nhất
- Trang Web Cá độ Bóng đá Của Việt Nam
- Casino En Ligne Avis
- Casino En Ligne France
- Casino En Ligne
- 꽁머니 토토
- Casino Online Non Aams
- Migliori Casino Non AAMS
- Meilleur Casino En Ligne
- Casino En Ligne France Légal
- Casino En Ligne France Légal
- Casinos En Ligne